The Waiting Time Paradox, or, Why Is My Bus Always Late?
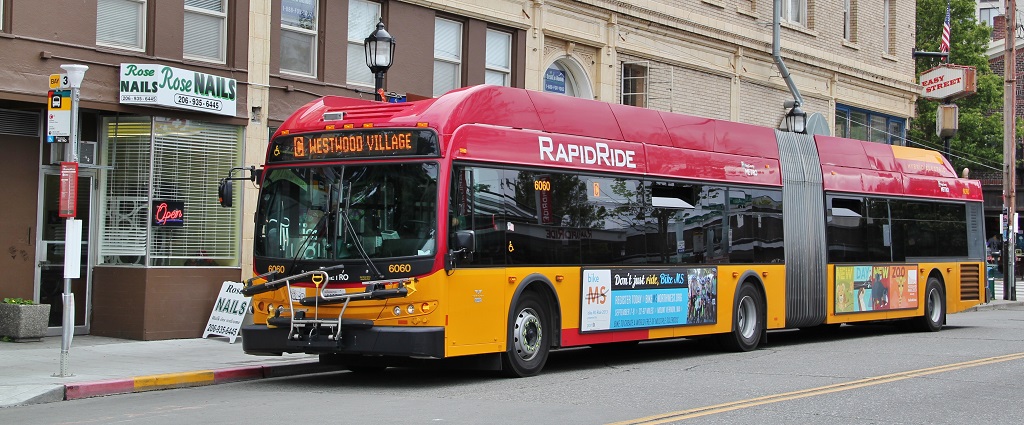 Image Source: Wikipedia License CC-BY-SA 3.0
Image Source: Wikipedia License CC-BY-SA 3.0
If you, like me, frequently commute via public transit, you may be familiar with the following situation:
You arrive at the bus stop, ready to catch your bus: a line that advertises arrivals every 10 minutes. You glance at your watch and note the time... and when the bus finally comes 11 minutes later, you wonder why you always seem to be so unlucky.
Naïvely, you might expect that if buses are coming every 10 minutes and you arrive at a random time, your average wait would be something like 5 minutes. In reality, though, buses do not arrive exactly on schedule, and so you might wait longer. It turns out that under some reasonable assumptions, you can reach a startling conclusion:
When waiting for a bus that comes on average every 10 minutes, your average waiting time will be 10 minutes.
This is what is sometimes known as the waiting time paradox.
I've encountered this idea before, and always wondered whether it is actually true... how well do those "reasonable assumptions" match reality? This post will explore the waiting time paradox from the standpoint of both simulation and probabilistic arguments, and then take a look at some real bus arrival time data from the city of Seattle to (hopefully) settle the paradox once and for all.
![[img: Chutes and Ladders animated simulation]](http://jakevdp.github.io/images/ChutesAndLadders-sim.gif)